The argument that altruism in sterile ants which work for the benefit of their community can be explained by kin selection has been overturned by mathematical analysis by Martin Nowak, author of Evolutionary Dynamics, in a paper published in Nature. Instead, altruism turns out to be a thoroughly naturally selected affair. Read more here.
|
THING ONE:
The argument that altruism in sterile ants which work for the benefit of their community can be explained by kin selection has been overturned by mathematical analysis by Martin Nowak, author of Evolutionary Dynamics, in a paper published in Nature. Instead, altruism turns out to be a thoroughly naturally selected affair. Read more here.
1 Comment
Nayab Abidi's essay "Mitochondria to determine species: An Insight into DNA Bar Codes" is now up in the Biology section, bringing the Biology essay count to 4, equalling those of Physics. We are proud we have finally caught up and we hope you are too. Here is an excerpt:
Researchers believe that this particular sequence becomes unique after two species split from their common ancestor. However, researchers will be researchers. Molecular biologist Dan Mishmar and his colleagues at the Ben-Gurion University of the Negev in Beer-Sheva, Israel, put forward this new hypothesis; that the bar codes do not become unique just ‘because’ the two species split but they might even be important ‘drivers’ of the process of speciation. What they propose is that this particular stretch of mitochondrial DNA might be responsible for undermining the reproductive compatibility within a species when it conflicts with the sequences in the nucleus. Once again, comments are welcome on this blog post. We hope to make the Work Section even better now that it has completely gotten off the ground. If you have any suggestions of topics you feel could create a fantabulous essay, do let us know. Furthermore, we are now again looking for new contributors: bloggers, essayists, artists, and even film-makers. If you want to join our team, navigate to the Contact tab and let us know! Short video appealing for flood relief. More short videos in the future. Mehr
The top fifty physics students of Pakistan, just back from their high profile tour of Pakistan's nuclear reactor near Islamabad, sat in a small auditorium listening intently to one engaging speaker.
I learnt something very interesting that day. While the government would love more students to go into nuclear physics, what they needed now were students who would model the weather. Pakistan, along with everyone else, was going to be hit by the results of global warming and it was imperative for us to be prepared. But that wasn't the scary bit. The United States was actually spending a lot to model the weather of many parts of the globe, including our neighbourhood. When our scientists asked for data and simulations the request was denied. Weather modelling was being considered as a potential weapon and therefore all critical information was highly classified. I used to play Red Alert 2 on my PC. The Allied weather storm was as powerful, if not more deadly than the Soviet nuclear missile. But it did not come with bad press, or the curse of deformed children. It was infact untraceable to its source. I am in no way downplaying the role of global warming. It has made all of this much more dangerous. Warmer waters means more violent storms and much heavier rains. America itself has been hit by the worse storms in decades. Weather is a funny thing because of the Butterfly Effect. It follows no given pattern and is in essence unpredictable. But our minds have grown sharper and our computing power stronger. We already seed clouds to cause rains. There may come a time when we can cause massive storms by just a small change in the conditions which will give the system a nudge to its more damaging state. I don't think we are there yet. Though some theorists (Zaid Hamid) certainly believe that our enemies have caused these very floods. And while you may not be a fan he does have something unique to say about most things. No conventional war could have brought this much devastation. But maybe we just got bombed. Hassan Photoblogs describe the flooding in different ways. The devastation wrought only goes to show how there is an overwhelming need not just for immediate resource-provision but relief efforts that are invested in over a period of many years, efforts that can help rehabilitate and rebuild the lives of the affectees. It also demonstrates the need to recognize how disastrous slow responses to a calamity can be, especially a calamity which shows no end in sight. Here are some of the pictures printed by different news agencies around the world. Captions have been placed as they appeared in print.
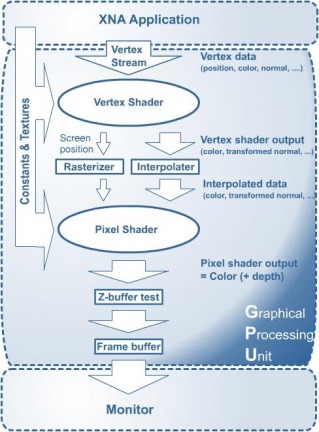
Computer gamers often do not appreciate the amount of effort that goes into making a single commercial video game. Making a game is one of the most sophisticated forms of software engineering, with the coders tweaking the very basic hardware configurations in order to achieve real-time rendering and response that is required of every game. Computer graphics have, in recent times taken an exponential leap, and this is reflected in any Pixar film we see on the telly. However, what is displayed on the screen in a pre-rendered film, the product of probably months of computer time. In a game, however, what you see on the screen is a direct consequence of your input to the machine, which means that the image on screen has to be generated there and then. To do this seamlessly and quickly requires a potent combination of theoretical linear algebra along with complex bit packing and explicit memory management, not just on your CPU, but also on your GPU. The “rendering pipeline”, as it is called, comprises of all the stages starting from the input data to your graphics card, and displaying the final output on the screen. Observe the following diagram of the pipeline: I came across this diagram while following Riemers Grootjans’s excellent tutorial on XNA Game Studio, an add-on to Microsoft Visual Studio and the newest “shell” for DirectX. The application we create in XNA passes a series of “vertices” to the GPU as a “vertex stream”. XNA mainly deals with outputting geometric primitives like triangles to the screen, and each polygon has multiple vertices. Each vertex comprises of a 3D position and other optional properties like a Color, a Normal, a Texture coordinate, etc. All these vertices are then simply passed on to the GPU, and it takes care of the rest. However, upon opening up the GPU, we find that there are multiple components to it, and what kind of computer scientists would we be if we didn’t throw away other people’s techniques and develop our own? Most of the GPU work used to be performed by fixed-function hardware in the past, but nowadays, each component is much more programmable in languages such as High Level Shader Language. I will now attempt to explain some of the more interesting parts of the GPU.
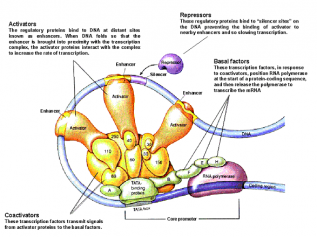
The vertex shader is the component that receives the initial vertex stream being passed to the GPU. This does the initial processing on the raw stream, and determines what the user wants drawn by the many vertices. For example, using 6 vertices, you could draw a hexagon, 2 triangles, or anything else. The “what to draw” information is encoded in something that is passed to the GPU even before the vertex stream. This is called the Vertex Declaration, and it tells the graphics card something like, “OK, you’re going to be receiving a stream of vertices, and you have to draw at most 3 triangles from it.” The vertex shader then passes the declaration and the required vertices to two things: the Rasterizer, and the Interpolator, which filter and process the data, and pass on the required information to the Pixel Shader. Both the Rasterizer and the Interpolator pass their data on to the most fun part of the GPU, the Pixel Shader. It is the job of the pixel shader to determine the color of every pixel on screen, nothing more nothing less. First, the data from the Rasterizer acts like a sieve. The rasterizer tells us exactly which pixels we need to be concerned with, and so we throw away the rest. The data from the interpolator is then used to determine the color of individual pixels on screen, but because the pixel shader is now programmable, this may not always be the case. The pixel shader is like a programmable function, which leaves it’s effects as creative as the programmer. I recently tweaked the pixel shader to make all the graphics appear in sepia mode. The pixel shader acts like a final filter between your data and what is displayed on screen, and a lot of cool effects can be implemented. Now someone might ask, “Why don’t we filter the colors before passing them to the GPU? Wouldn’t that be simpler?” The answer is this: The GPU is so much better at it than the CPU. Today, most high-end graphics cards have dedicated hardware, RAM and processor. This hardware is meant for rendering purposes, so why not use it? The CPU has a dozen other things it needs to do, like run the physics engine, check for collision, process input, gather network messages, and much more. Taking advantage of the parallelism between your CPU and GPU can go a long way to boost performance, especially in a situation where you’re forced to code in straight C (nightmare) just so your code runs faster. Anyways, just a heads-up for all Graphics and Vision enthusiasts. This is all very cool stuff, but it’s slowly becoming more and more subtle and complex for the much-required performance boosts. Anyway, until next time! Asadullah  Can Pakistan be Governed -The New York TImes Note: This is a political blog post. Views are not the general opinion of the blog but only of the author. I don't normally talk about politics, but all of a sudden I feel an urge to. We as a people are currently blaming Zardari for most governmental inefficiencies, problems etc. What we must never forget is that he is in a democratic office. All decisions are run through Parliament and they all are passed with majorities and little opposition. We must not forget that the system supports and encourages all decisions. While we may choose to ridicule Zardari and wish him to be dead, he is just one man. The system wants you to focus your anger and hate on this one man and spare the system itself. This man will be replaced. Do not then be surprised that things haven't gotten better. It is not single men that rule countries but the echelons of power behind them. When they change will the real change come. Do not forget that. In other news, my maid started a literacy class at her home. What started with 35 enthusiastic other maids and their daughters has now been reduced to 6 odd people because the men don't want them to get any sort of an education. She is facing a divorce as her husband is having a fling with another lady. Science will save us all. Hassan All cells contain the same amount of information for performing their function. This information is available in the form of genes. One may then wonder how is it that all cells contain the same genes yet a variety of them exist in the form of muscle cells, liver cells, epithelial cells etc. Cells use information or signals from their surroundings to turn on certain genes within a particular cell whereas other genes remain switched off. So, for example, the insulin producing gene would be switched off in a liver cell whereas it would be functional in a pancreatic beta-cell. These external factors activate gene expression by the coordinated action of trans-acting transcriptional factors that bind to cis-acting regulatory elements such as promoters, enhancers, etc. Promoters are regions of DNA usually found upstream of the gene that’s transcribed. Promoters contain specific DNA sequences and response elements that can bind transcription factors, which than recruit RNA polymerase II, an enzyme necessary for transcription to take place. Enhancers on the other hand may be found upstream or downstream or even within the gene they activate. Enhancers are DNA sequences often found thousands of base pairs away from the gene they transcribe. One may then ask how binding of proteins at enhancers regulate transcription of genes thousands of base pairs away. One explanation is that the enhancer does not act on the promoter region itself, but is bound by activating proteins that interact with the mediator complex which is required for transcription at the promoter, bending the DNA in a loop.
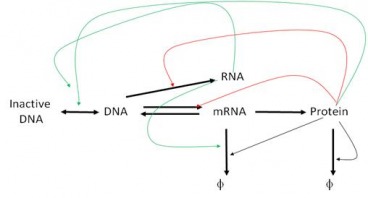
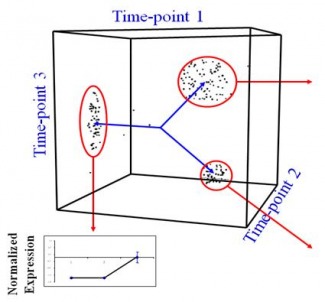
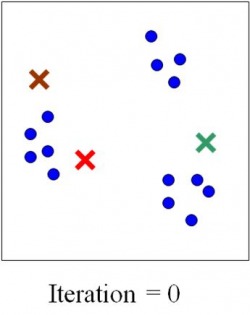
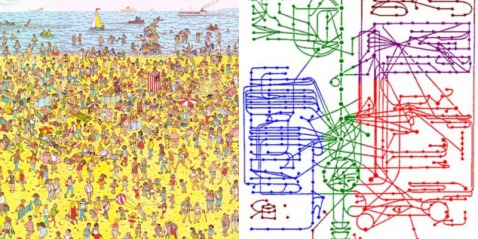
While the mechanisms by which promoters control gene transcription are well understood, the mechanism of regulation by enhancers has not been characterized. Kim et al tried to investigate this mechanism in a paper published in Nature in April 2010. For their purpose they identified 12,000 enhancers in activity-regulated mouse cortical neuronal cells. They showed that these enhancer sequences are involved in regulating gene expression during neuronal activity within the cells. However, the most interesting find of the paper has been that of another non-coding RNA, called enhancer RNA (eRNA). eRNA is an addition to the class of non-coding RNA’s that had previously been identified including tRNA (carries amino acids and helps in translation), microRNA, siRNA etc. Non-coding RNA’s are functional RNA’s that do not get translated into a protein. The authors identified enhancer sequences involved in activity-regulated gene expression. The membranes were depolarized using potassium chloride. 12,000 enhancers were identified by mapping enhancer bound CBP sites (p300/CBP is a co-transcriptional system). To exclude any uncharacterized promoters that might have been selected they further narrowed their search by selecting histone marks specific to enhancer regions: enhancers have high levels of mono-methylated histone 3 lysine 4 and low tri-methylated histone 3 lysine 4. In the paper they analyzed the binding of three transcriptional factors and RNA pol II to these enhancers. Kim et al found that 25% of the enhancers bound to RNA pol II, and an RNA transcript was found at each of these enhancers. This eRNA is less than 2kb in size, is transcribed bi-directionally and lacks the polyA chain found in mature RNA’s. Moreover, changes in eRNA related to changes in mRNA levels from nearby genes. For the case examined in the paper (that of Arc enhancers), it was found that in order for eRNA synthesis to take place a promoter is required. However, the binding of transcription factors and recruitment of RNA Pol is independent of the presence of a promoter. In addition, since only a fraction of CBP-bound enhancers produced eRNA, it was concluded that other factors besides CBP binding might be required for eRNA synthesis. What however, remains inconclusive is how this eRNA is involved in enhancer function. One strategy may be that eRNA’s are involved in activating the corresponding promoters as suggested by Bing Ren in his commentary on the paper ‘Enhancers make non-coding RNA’ in Nature. “eRNA synthesis may be necessary for activating the corresponding promoter. This possibility is supported by indirect evidence from the β-globin gene locus. When a 1-kilobase-long transcription termination sequence was inserted between the enhancer and the promoter sequences for the β-globin gene, trans cription from the enhancer ended prematurely, and activation of the promoter by the enhancer was significantly reduced. This observation suggests that enhancer transcription may be necessary for activating promoters.” Another possibility suggested by Kim et al is that the process of eRNA synthesis could be required to maintain chromatin at enhancers in a state necessary for enhancer function. This hypothesis however, can be confirmed by introducing a transcription stop element at enhancers, which would result in no eRNA being formed, so if they are required this would affect the transcriptional activation of targets by enhancers (the same could be done using RNAi). If these eRNA’s are indeed necessary for enhancer function, enhancers can be inhibited by interfering with the transcription of eRNA. Since enhancer sequences are involved in causing certain diseases, understanding their function can take us one step further to finding a cure for these illnesses. Anum The sequencing of the human genome was one of those events, those oft-criticized scientific frontiers which are controversial and scary and aren’t explained to the public very well. What they should have been told is that it offers the greatest amount of insight into modern medicine, and only now as Biology capitalizes on the advances made by the Human Genome Project is the effect apparent. The point of Systems Biology is to describe not individual genes and what proteins they make but to step away from the Central Dogma and treat it as a whole, with networks and pathways that can only be understood holistically. The whole apparently is greater than the sum of its parts. We know that genes can interact with each other – epistasis – and we know that genes create proteins that can come and affect the workings of DNA also. But the whole story is a great deal more complex than that, and it involves all sorts of players – mRNA, chromatin, micro RNA. In fact, if we modify the central dogma to depict current research, it would look something like this: One idea is to be able to describe not just genes, but whole sets of genes and see what distinguishing features of such sets are. Let’s say that we can take all the genes in the genome and see how they are transcribed into mRNA and then we can plot the expression level of each gene (measured using microarrays or DNA chips) on a graph against time. If we make such a graph, certain patterns begin to emerge – let’s say genes M, X, and Z for instance show a periodic pattern with a peak in the S-G2 phase of the cell cycle. Now let’s clump together all the genes that show such similar patterns. If we do this over various time points – 0, 10 minutes, 20 minutes and so on – we can build up a huge graph where we can put each gene based on its expression level at the different time points. What we get is a really awesome multiple-dimension graph called a time-dependent expression profile that looks like the diagram below. Here, the gene is being plotted based on its position in 3 time points – so if a gene has A expression level at time 1, B at time 2 and C and time 3 then its coordinate will be (A, B, C) Tavazoie et al in a seminal paper in Nature Genetics way back when in 1999 came up with an algorithm called the K-means algorithm that could clump groups of genes together. The idea is the same – genes closer together are more likely to be part of a group because they show a similar expression pattern. What they did was come up with an unsupervised algorithm. First, the experimenter gives a guess for the number of clusters or sets of genes – this guess is the value of K. Let’s say I thought that this has three clusters (K=3) as is fairly obvious: Once the guess has been given, the algorithm defines random positions for the ‘centroids’(the cross on the diagram) and then iteratively tries to get the centroids into the center of the gene sets, trying to minimize the distance between the genes and their closest centroid. The number of centroids is the number K. This algorithm is limited by the ‘guess’ we give it so of course ideally we would like an algorithm that doesn’t need a guess to get the right number of sets. This is the basis of newer algorithms which are called hierarchical clustering algorithms. Once the iterations are complete, we are given an output for the number of clusters or the real value of K. Now, what similarity could these genes in a cluster possibly have? As it turns out, they have functional similarities so genes which make ribosomes for example belong to a cluster. But can we ask an even deeper question? Do these genes have specific sequences that allow them to be regulated in the same manner? Yes of course they do, and we can find out by looking for certain sequences called ‘motifs’ upstream of each gene in a cluster. If we find sequences that seem to be appearing more than is possible through randomness, the sequence is said to be over-represented and it could be a motif. All this analysis can tell us in a system-specific manner how genes are part of networks called ‘regulons’. Could a cluster of genes made for say, organizing the mitochondria, be interacting with another cluster of genes which make transport proteins? This can be imagined by zooming out of the picture. Of course, it could also be thought of as a Where’s Waldo game. Compare these two images below – which one seems more complex? This brings me to the point in the beginning about the prospects for modern medicine. If we know how these sets of genes interact, we could in principle map all sorts of functional networks, the same way we do for complex metabolic cascades. Now imagine tinkering with the individual DNA of a number of people, all with a rudimentary system of interactions that we know about. Assuming that we know where in this system each person differs from the others, imagine the possibilities – medicine specific to that person as opposed to a generic applies-to-all. Furthermore, as we understand more principles inherent to such networks, we’ll be able to delve in into questions that deal with the genome as a whole, as opposed to isolated effects in isolated places. That’s the beauty of it – you’re making sense by looking at the larger picture not by going down to the molecular level. Kamil
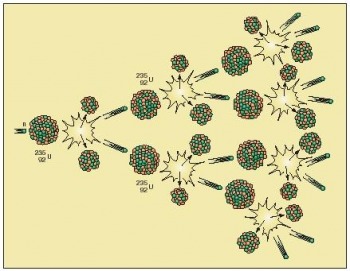
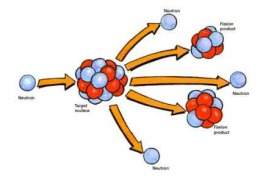 It’s a summer Saturday. I’m here in LUMS for a Math department research project but free for the weekend. Before going to sleep last night, I had planned to go to my sister’s place at Allama Iqbal Medical College, near Jinnah Hospital - quite far from LUMS. Now I am up, have nothing to do and am still in bed. I should start moving now, but I look outside the window, the sun is shining brightly and it’s really hot. I imagine myself getting up, picking up my bags, going all the way to the main gate, hiring a tuk-tuk (rickshaw), waiting on the signals and sweating all around. I don’t want to go. No, I want to meet my sister. Okay so, I want to go but I don’t want to travel. That's pretty abnormal. Wait a second - that’s not completely abnormal! While trying to get up, I have this idea to develop some technique that could convert my mass into energy so I could travel with the speed of light, reach my sister’s place and convert myself back into matter. Let me start with an explanation of the famous E = mc2 thing. It says that the total energy of a body at rest is equal to the product of its rest mass m and a suitable conversion factor to transform from units of mass to units of energy. In other words, this equation says that matter and energy are convertible. This conversion actually happens! Nuclear fission is a nuclear reaction in which the nucleus of an atom splits into lighter nuclei often producing free neutrons and protons (in the form of gamma rays). Such splitting of an atom releases an incredible amount of heat and gamma radiation, or radiation made of high-energy photons. The two atoms that result from the fission later release beta radiation (super fast electrons) and gamma radiation of their own as well. The energy released by a single fission comes from the fact that the fission products and the neutrons, together, weigh less than the original U-235 atom. The difference in weight is converted directly to energy at a rate governed by the equation E = mc2. Practically of course, in a nuclear fission reaction, 7% of the matter is converted into energy. Conversely, in the phenomenon of pair production, two gamma rays give an electron-positron pair essentially converting energy into mass. It refers to the creation of an elementary particle and its antiparticle, usually from a photon (or another neutral boson). This is allowed, provided there is enough energy available to create the pair – at least the total rest mass energy of the two particles and that the situation allows both energy and momentum to be conserved. When we talk about converting energy into matter we cannot just create matter out of energy. There are various laws of conservation that have to be followed. Whenever we try to convert energy into matter, it gives a matter and anti-matter pair which immediately combines back to energy. So it takes a huge amount of energy to keep them from combining. In the LHC at CERN, high speed protons are colliding to produce energy just as it happened before the existence of the universe. Protons are racing around a 27-kilometer ring at nearly the speed of light colliding billions of times per second. If we get the results of that experiment, that would be an insight to many phenomenon of the early universe and will explain a great deal about the mass-energy conversion thing. It would take 300 thousand times the age of the Universe for the LHC to produce one kilogram of mass. Still, it does produce mass from energy! The problem is that the mass particles that are produced decay into smaller components which in turn decay into smaller ones until all that remains are stable particles such as those that constitute the atoms of matter we see all around – protons and neutrons - only dispersed. We can also count neutrinos and electrons, but those are much less massive than protons and neutrons. Once we convert matter into energy, the matter is destroyed. And it becomes highly improbable to recollect that energy and convert it back into matter anti-matter pair. With these facts and taking into account all the experimental and technical difficulties, it becomes extremely ridiculous to think that some day we would actually convert ourselves into energy and control ourselves then. So yes, we will have to wait till we can govern some system to convert effectively all our particles into energy and more importantly devise some plan to preserve that energy, making it unique, transport it and reconvert it back into matter whenever we want. Let’s just take comfort in the fact that many great inventions start from ideas that arise from laziness! Rabia |
The Box Move blog is no longer active since the founding team has graduated. The archives will remain online.
AFFILIATES: 
The SPROJ Forum for the SSE 2012 batch. Discuss potential Senior Projects here.

Brain Talk is an online resource and forum for all things Psychology and Neuroscience.
MUSIC FOR GEEKS:
Featured: Art Tatum Three Letter Word ON THE FRINGE:
Learn something
The story of a how a YouTube video of a blind man biking down a mountain inspired good non-fiction writing on echolocation. You may find it useful for your own writing. Read here. An awful waste of space?
Amidst NASA's budgetary cuts and scientists' renewed vigor in justifying Space Programs, it is important to shed some light on the background. Click here for a succinct overview of the Search for Extra-Terrestrial Intelligence(SETI) project. Manto ka Muqaddama
Pakistaniat.com publishes, on the anniversary of Saadat Hasan Manto's death, a sampling of his works, a tribute to him as well as articles chronicling the obscenity trial he was tried for. Read all three parts of the series here. Not Another 2010 List
The folks over at The Last Word put up their list for the best non-fiction in the past year, including 'The Mind's Eye' which is very hard to find in bookstores, indeed! Read the full list here. Bird Conspiracies
Leslie Kaufman at The New York Times tells us exactly why those birds flying above are dropping to their deaths. Click here to read the article. Spontaneous Solar Growth!
Reported at MadScience, scientists at MIT have found a way to create solar cells that can regenerate themselves like living organisms. Read more here. Lessons from Chernobyl
Decades after the radiation disaster at Chernobyl, scientists elucidate how plant life has been thriving in the highly radioactive environment. Read more here. Rolling Ribbons
MIT Scientists revisit Galileo's famous inclined plane experiment, this time with polymer ribbons and discover complex results. Read here. A Lifetime, Washed Away
Pakistani author Daniyal Mueenuddin writes in the NY Times about the aftermath of the flood and displaced people. Read more here of the article posted by 3QD. On String theory and Materials Science
Click here to find out how physicists at MIT are using ideas of gauge/gravity duality to explain properties of superconductors. That's why you're irrational!
Newsweek's Sharon Begley provides a fascinating argument for why evolution may favor irrationality. I particularly liked the examples she picked. Read here. Just when you wanted a gene kit
The US Food and Drug Administration held hearings n the 19th and 20th of July to talk about the validity of tests which were sold directly to the public which gives consumers direct access to to their genomes. Should it be regulated? Read more here. On Trees and Prisons
In a 6 minute talk on Ted.com, Nalini Nadkarni (shown above) talks about her ideas of incorporating conservation into prison programs. Watch the talk and read Nadkarni's fascinating biography here. These Lungs are made in USA
Stem Cell Biology takes huge leaps forward with the new advances made in lung transplants based on using the lungs extracellular architecture. Read more from Nature here. Economist Special Reports
Ten years after Craig Venter revealed the first working draft of the entire Human Genome, this special report demonstrates how Biology is now at the brink of something brilliant - just recently, the draft of the entire genome of the Neanderthal was revealed. Suck on that, sceptics! Orbit Stories
Bobby Satcher, astronaut, the first orthopedic surgeon in space. Read all about his tales here on MITnews. 
Craig Venter Creations
Researchers create world's first fully synthetic self replicating, living cell. Massive fuss about limitless monster potential possible. Read the NewScientist article here. Watch the Ted.com talk by Craig Venter here. TAGS
All
FEEDS:
ARCHIVES
January 2013
|
||||||






 RSS Feed
RSS Feed

















