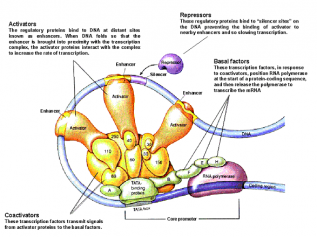
While the mechanisms by which promoters control gene transcription are well understood, the mechanism of regulation by enhancers has not been characterized. Kim et al tried to investigate this mechanism in a paper published in Nature in April 2010. For their purpose they identified 12,000 enhancers in activity-regulated mouse cortical neuronal cells. They showed that these enhancer sequences are involved in regulating gene expression during neuronal activity within the cells. However, the most interesting find of the paper has been that of another non-coding RNA, called enhancer RNA (eRNA). eRNA is an addition to the class of non-coding RNA’s that had previously been identified including tRNA (carries amino acids and helps in translation), microRNA, siRNA etc. Non-coding RNA’s are functional RNA’s that do not get translated into a protein.
The authors identified enhancer sequences involved in activity-regulated gene expression. The membranes were depolarized using potassium chloride. 12,000 enhancers were identified by mapping enhancer bound CBP sites (p300/CBP is a co-transcriptional system). To exclude any uncharacterized promoters that might have been selected they further narrowed their search by selecting histone marks specific to enhancer regions: enhancers have high levels of mono-methylated histone 3 lysine 4 and low tri-methylated histone 3 lysine 4. In the paper they analyzed the binding of three transcriptional factors and RNA pol II to these enhancers. Kim et al found that 25% of the enhancers bound to RNA pol II, and an RNA transcript was found at each of these enhancers. This eRNA is less than 2kb in size, is transcribed bi-directionally and lacks the polyA chain found in mature RNA’s. Moreover, changes in eRNA related to changes in mRNA levels from nearby genes. For the case examined in the paper (that of Arc enhancers), it was found that in order for eRNA synthesis to take place a promoter is required. However, the binding of transcription factors and recruitment of RNA Pol is independent of the presence of a promoter. In addition, since only a fraction of CBP-bound enhancers produced eRNA, it was concluded that other factors besides CBP binding might be required for eRNA synthesis.
What however, remains inconclusive is how this eRNA is involved in enhancer function. One strategy may be that eRNA’s are involved in activating the corresponding promoters as suggested by Bing Ren in his commentary on the paper ‘Enhancers make non-coding RNA’ in Nature.
“eRNA synthesis may be necessary for activating the corresponding promoter. This possibility is supported by indirect evidence from the β-globin gene locus. When a 1-kilobase-long transcription termination sequence was inserted between the enhancer and the promoter sequences for the β-globin gene, trans cription from the enhancer ended prematurely, and activation of the promoter by the enhancer was significantly reduced. This observation suggests that enhancer transcription may be necessary for activating promoters.”
Another possibility suggested by Kim et al is that the process of eRNA synthesis could be required to maintain chromatin at enhancers in a state necessary for enhancer function. This hypothesis however, can be confirmed by introducing a transcription stop element at enhancers, which would result in no eRNA being formed, so if they are required this would affect the transcriptional activation of targets by enhancers (the same could be done using RNAi).
If these eRNA’s are indeed necessary for enhancer function, enhancers can be inhibited by interfering with the transcription of eRNA. Since enhancer sequences are involved in causing certain diseases, understanding their function can take us one step further to finding a cure for these illnesses.
Anum
 RSS Feed
RSS Feed