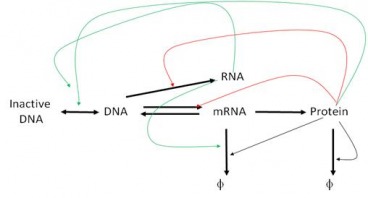
The point of Systems Biology is to describe not individual genes and what proteins they make but to step away from the Central Dogma and treat it as a whole, with networks and pathways that can only be understood holistically. The whole apparently is greater than the sum of its parts. We know that genes can interact with each other – epistasis – and we know that genes create proteins that can come and affect the workings of DNA also. But the whole story is a great deal more complex than that, and it involves all sorts of players – mRNA, chromatin, micro RNA. In fact, if we modify the central dogma to depict current research, it would look something like this:
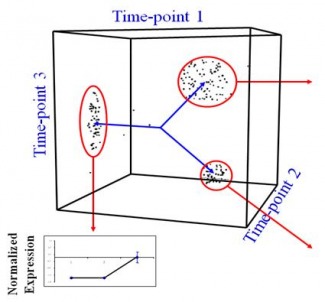
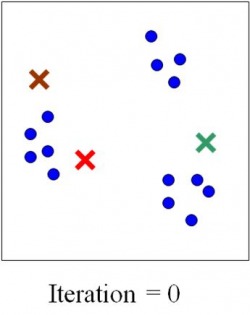
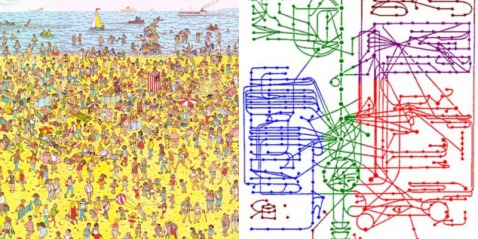
Once the iterations are complete, we are given an output for the number of clusters or the real value of K. Now, what similarity could these genes in a cluster possibly have? As it turns out, they have functional similarities so genes which make ribosomes for example belong to a cluster. But can we ask an even deeper question? Do these genes have specific sequences that allow them to be regulated in the same manner? Yes of course they do, and we can find out by looking for certain sequences called ‘motifs’ upstream of each gene in a cluster. If we find sequences that seem to be appearing more than is possible through randomness, the sequence is said to be over-represented and it could be a motif. All this analysis can tell us in a system-specific manner how genes are part of networks called ‘regulons’. Could a cluster of genes made for say, organizing the mitochondria, be interacting with another cluster of genes which make transport proteins? This can be imagined by zooming out of the picture. Of course, it could also be thought of as a Where’s Waldo game. Compare these two images below – which one seems more complex?
Kamil
| tavazoie99.pdf |
 RSS Feed
RSS Feed